Identifying experiments and sessions of interest using the data manifest¶
This Jupyter notebook illustrates what data is available as part of the Visual Behavior - 2P dataset, and helps you to understand the experimental design and dimensions of the dataset. The notebook will demonstrate how to identify experiments and sessions that you may be interested in analyzing using the data manifests provided by the VisualBehaviorOphysProjectCache, and exploring the metadata columns that describe the experimental conditions including transgenic lines, targeted areas, imaging depths, microscopes that were used, session types, and dataset variants.
We will first install allensdk into your environment by running the appropriate commands below.
Install AllenSDK into your local environment¶
You can install AllenSDK locally with:
!pip install allensdk
Requirement already satisfied: allensdk in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (2.16.2) Requirement already satisfied: psycopg2-binary in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.9.9) Requirement already satisfied: hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.* in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.11.0) Requirement already satisfied: h5py in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.10.0) Requirement already satisfied: matplotlib in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.7.4) Requirement already satisfied: numpy<1.24 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.23.5) Requirement already satisfied: pandas==1.5.3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.5.3) Requirement already satisfied: jinja2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.1.2) Requirement already satisfied: scipy<1.11 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.10.1) Requirement already satisfied: six in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.16.0) Requirement already satisfied: pynrrd in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.0.0) Requirement already satisfied: future in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.18.3) Requirement already satisfied: requests in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.31.0) Requirement already satisfied: requests-toolbelt in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.0.0) Requirement already satisfied: simplejson in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.19.2) Requirement already satisfied: scikit-image in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.21.0) Requirement already satisfied: scikit-build in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.17.6) Requirement already satisfied: statsmodels in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.14.0) Requirement already satisfied: simpleitk in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.3.1) Requirement already satisfied: argschema in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.0.4) Requirement already satisfied: glymur in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.12.9.post1) Requirement already satisfied: xarray<2023.2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2023.1.0) Requirement already satisfied: pynwb in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.5.0) Requirement already satisfied: tables in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.8.0) Requirement already satisfied: seaborn in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.13.0) Requirement already satisfied: aiohttp in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.9.1) Requirement already satisfied: nest-asyncio in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.5.8) Requirement already satisfied: tqdm in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (4.66.1) Requirement already satisfied: ndx-events in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.2.0) Requirement already satisfied: boto3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.33.4) Requirement already satisfied: semver in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.0.2) Requirement already satisfied: cachetools in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (5.3.2) Requirement already satisfied: sqlalchemy in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.0.23) Requirement already satisfied: python-dateutil in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.8.2) Requirement already satisfied: pytz>=2020.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pandas==1.5.3->allensdk) (2023.3.post1) Requirement already satisfied: importlib-resources in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (6.1.1) Requirement already satisfied: jsonschema>=2.6.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (4.20.0) Requirement already satisfied: ruamel-yaml>=0.16 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.18.5) Requirement already satisfied: packaging>=21.3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from xarray<2023.2.0->allensdk) (23.2) Requirement already satisfied: attrs>=17.3.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (23.1.0) Requirement already satisfied: multidict<7.0,>=4.5 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (6.0.4) Requirement already satisfied: yarl<2.0,>=1.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.9.3) Requirement already satisfied: frozenlist>=1.1.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.4.0) Requirement already satisfied: aiosignal>=1.1.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.3.1) Requirement already satisfied: async-timeout<5.0,>=4.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (4.0.3) Requirement already satisfied: marshmallow<4.0,>=3.0.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from argschema->allensdk) (3.20.1) Requirement already satisfied: pyyaml in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from argschema->allensdk) (6.0.1) Requirement already satisfied: botocore<1.34.0,>=1.33.4 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (1.33.4) Requirement already satisfied: jmespath<2.0.0,>=0.7.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (1.0.1) Requirement already satisfied: s3transfer<0.9.0,>=0.8.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (0.8.2) Requirement already satisfied: lxml in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from glymur->allensdk) (4.9.3) Requirement already satisfied: MarkupSafe>=2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jinja2->allensdk) (2.1.3) Requirement already satisfied: contourpy>=1.0.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (1.1.1) Requirement already satisfied: cycler>=0.10 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (0.12.1) Requirement already satisfied: fonttools>=4.22.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (4.45.1) Requirement already satisfied: kiwisolver>=1.0.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (1.4.5) Requirement already satisfied: pillow>=6.2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (10.1.0) Requirement already satisfied: pyparsing>=2.3.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (3.1.1) Requirement already satisfied: setuptools in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynwb->allensdk) (56.0.0) Requirement already satisfied: nptyping in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynrrd->allensdk) (2.5.0) Requirement already satisfied: typing-extensions in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynrrd->allensdk) (4.8.0) Requirement already satisfied: charset-normalizer<4,>=2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (3.3.2) Requirement already satisfied: idna<4,>=2.5 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (3.6) Requirement already satisfied: urllib3<3,>=1.21.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (1.26.18) Requirement already satisfied: certifi>=2017.4.17 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (2023.11.17) Requirement already satisfied: distro in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (1.8.0) Requirement already satisfied: tomli in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (2.0.1) Requirement already satisfied: wheel>=0.32.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (0.42.0) Requirement already satisfied: networkx>=2.8 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (3.1) Requirement already satisfied: imageio>=2.27 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (2.33.0) Requirement already satisfied: tifffile>=2022.8.12 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (2023.7.10) Requirement already satisfied: PyWavelets>=1.1.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (1.4.1) Requirement already satisfied: lazy_loader>=0.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (0.3) Requirement already satisfied: greenlet!=0.4.17 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from sqlalchemy->allensdk) (3.0.1) Requirement already satisfied: patsy>=0.5.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from statsmodels->allensdk) (0.5.3) Requirement already satisfied: cython>=0.29.21 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (3.0.6) Requirement already satisfied: numexpr>=2.6.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (2.8.6) Requirement already satisfied: blosc2~=2.0.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (2.0.0) Requirement already satisfied: py-cpuinfo in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (9.0.0) Requirement already satisfied: msgpack in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from blosc2~=2.0.0->tables->allensdk) (1.0.7) Requirement already satisfied: zipp>=3.1.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from importlib-resources->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (3.17.0) Requirement already satisfied: jsonschema-specifications>=2023.03.6 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (2023.11.1) Requirement already satisfied: pkgutil-resolve-name>=1.3.10 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (1.3.10) Requirement already satisfied: referencing>=0.28.4 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.31.1) Requirement already satisfied: rpds-py>=0.7.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.13.2) Requirement already satisfied: ruamel.yaml.clib>=0.2.7 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from ruamel-yaml>=0.16->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.2.8) [notice] A new release of pip is available: 23.0.1 -> 23.3.1 [notice] To update, run: pip install --upgrade pip
Install AllenSDK into your notebook environment (good for Google Colab)¶
You can install AllenSDK into your notebook environment by executing the cell below.
If using Google Colab, click on the RESTART RUNTIME button that appears at the end of the output when this cell is complete,. Note that running this cell will produce a long list of outputs and some error messages. Clicking RESTART RUNTIME at the end will resolve these issues. You can minimize the cell after you are done to hide the output.
!pip install --upgrade pip
!pip install allensdk
Requirement already satisfied: pip in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (23.0.1) Collecting pip Downloading pip-23.3.1-py3-none-any.whl.metadata (3.5 kB) Downloading pip-23.3.1-py3-none-any.whl (2.1 MB) ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ 2.1/2.1 MB 42.3 MB/s eta 0:00:00 Installing collected packages: pip Attempting uninstall: pip Found existing installation: pip 23.0.1 Uninstalling pip-23.0.1: Successfully uninstalled pip-23.0.1 Successfully installed pip-23.3.1 Requirement already satisfied: allensdk in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (2.16.2) Requirement already satisfied: psycopg2-binary in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.9.9) Requirement already satisfied: hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.* in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.11.0) Requirement already satisfied: h5py in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.10.0) Requirement already satisfied: matplotlib in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.7.4) Requirement already satisfied: numpy<1.24 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.23.5) Requirement already satisfied: pandas==1.5.3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.5.3) Requirement already satisfied: jinja2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.1.2) Requirement already satisfied: scipy<1.11 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.10.1) Requirement already satisfied: six in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.16.0) Requirement already satisfied: pynrrd in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.0.0) Requirement already satisfied: future in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.18.3) Requirement already satisfied: requests in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.31.0) Requirement already satisfied: requests-toolbelt in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.0.0) Requirement already satisfied: simplejson in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.19.2) Requirement already satisfied: scikit-image in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.21.0) Requirement already satisfied: scikit-build in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.17.6) Requirement already satisfied: statsmodels in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.14.0) Requirement already satisfied: simpleitk in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.3.1) Requirement already satisfied: argschema in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.0.4) Requirement already satisfied: glymur in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.12.9.post1) Requirement already satisfied: xarray<2023.2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2023.1.0) Requirement already satisfied: pynwb in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.5.0) Requirement already satisfied: tables in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.8.0) Requirement already satisfied: seaborn in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.13.0) Requirement already satisfied: aiohttp in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.9.1) Requirement already satisfied: nest-asyncio in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.5.8) Requirement already satisfied: tqdm in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (4.66.1) Requirement already satisfied: ndx-events in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (0.2.0) Requirement already satisfied: boto3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (1.33.4) Requirement already satisfied: semver in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (3.0.2) Requirement already satisfied: cachetools in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (5.3.2) Requirement already satisfied: sqlalchemy in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.0.23) Requirement already satisfied: python-dateutil in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from allensdk) (2.8.2) Requirement already satisfied: pytz>=2020.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pandas==1.5.3->allensdk) (2023.3.post1) Requirement already satisfied: importlib-resources in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (6.1.1) Requirement already satisfied: jsonschema>=2.6.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (4.20.0) Requirement already satisfied: ruamel-yaml>=0.16 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.18.5) Requirement already satisfied: packaging>=21.3 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from xarray<2023.2.0->allensdk) (23.2) Requirement already satisfied: attrs>=17.3.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (23.1.0) Requirement already satisfied: multidict<7.0,>=4.5 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (6.0.4) Requirement already satisfied: yarl<2.0,>=1.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.9.3) Requirement already satisfied: frozenlist>=1.1.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.4.0) Requirement already satisfied: aiosignal>=1.1.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (1.3.1) Requirement already satisfied: async-timeout<5.0,>=4.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from aiohttp->allensdk) (4.0.3) Requirement already satisfied: marshmallow<4.0,>=3.0.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from argschema->allensdk) (3.20.1) Requirement already satisfied: pyyaml in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from argschema->allensdk) (6.0.1) Requirement already satisfied: botocore<1.34.0,>=1.33.4 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (1.33.4) Requirement already satisfied: jmespath<2.0.0,>=0.7.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (1.0.1) Requirement already satisfied: s3transfer<0.9.0,>=0.8.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from boto3->allensdk) (0.8.2) Requirement already satisfied: lxml in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from glymur->allensdk) (4.9.3) Requirement already satisfied: MarkupSafe>=2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jinja2->allensdk) (2.1.3) Requirement already satisfied: contourpy>=1.0.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (1.1.1) Requirement already satisfied: cycler>=0.10 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (0.12.1) Requirement already satisfied: fonttools>=4.22.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (4.45.1) Requirement already satisfied: kiwisolver>=1.0.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (1.4.5) Requirement already satisfied: pillow>=6.2.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (10.1.0) Requirement already satisfied: pyparsing>=2.3.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from matplotlib->allensdk) (3.1.1) Requirement already satisfied: setuptools in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynwb->allensdk) (56.0.0) Requirement already satisfied: nptyping in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynrrd->allensdk) (2.5.0) Requirement already satisfied: typing-extensions in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from pynrrd->allensdk) (4.8.0) Requirement already satisfied: charset-normalizer<4,>=2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (3.3.2) Requirement already satisfied: idna<4,>=2.5 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (3.6) Requirement already satisfied: urllib3<3,>=1.21.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (1.26.18) Requirement already satisfied: certifi>=2017.4.17 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from requests->allensdk) (2023.11.17) Requirement already satisfied: distro in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (1.8.0) Requirement already satisfied: tomli in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (2.0.1) Requirement already satisfied: wheel>=0.32.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-build->allensdk) (0.42.0) Requirement already satisfied: networkx>=2.8 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (3.1) Requirement already satisfied: imageio>=2.27 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (2.33.0) Requirement already satisfied: tifffile>=2022.8.12 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (2023.7.10) Requirement already satisfied: PyWavelets>=1.1.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (1.4.1) Requirement already satisfied: lazy_loader>=0.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from scikit-image->allensdk) (0.3) Requirement already satisfied: greenlet!=0.4.17 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from sqlalchemy->allensdk) (3.0.1) Requirement already satisfied: patsy>=0.5.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from statsmodels->allensdk) (0.5.3) Requirement already satisfied: cython>=0.29.21 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (3.0.6) Requirement already satisfied: numexpr>=2.6.2 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (2.8.6) Requirement already satisfied: blosc2~=2.0.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (2.0.0) Requirement already satisfied: py-cpuinfo in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from tables->allensdk) (9.0.0) Requirement already satisfied: msgpack in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from blosc2~=2.0.0->tables->allensdk) (1.0.7) Requirement already satisfied: zipp>=3.1.0 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from importlib-resources->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (3.17.0) Requirement already satisfied: jsonschema-specifications>=2023.03.6 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (2023.11.1) Requirement already satisfied: pkgutil-resolve-name>=1.3.10 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (1.3.10) Requirement already satisfied: referencing>=0.28.4 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.31.1) Requirement already satisfied: rpds-py>=0.7.1 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from jsonschema>=2.6.0->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.13.2) Requirement already satisfied: ruamel.yaml.clib>=0.2.7 in /opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages (from ruamel-yaml>=0.16->hdmf!=3.5.*,!=3.6.*,!=3.7.*,!=3.8.*->allensdk) (0.2.8)
Import necessary packages¶
import numpy as np
from allensdk.brain_observatory.behavior.behavior_project_cache import VisualBehaviorOphysProjectCache
/opt/hostedtoolcache/Python/3.8.18/x64/lib/python3.8/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html from .autonotebook import tqdm as notebook_tqdm
First, load the project cache - your access point for all tables and data¶
# Update this to a valid directory in your filesystem
output_dir = r"\Data\visual_behavior_ophys_cache_dir"
cache = VisualBehaviorOphysProjectCache.from_s3_cache(cache_dir=output_dir)
/home/runner/work/AllenSDK/AllenSDK/allensdk/api/cloud_cache/cloud_cache.py:439: MissingLocalManifestWarning: This cache directory appears to contain data files, but it has no record of what those files are. You might want to consider running
VisualBehaviorOphysProjectCache.construct_local_manifest()
to avoid needlessly downloading duplicates of data files that did not change between data releases. NOTE: running this method will require hashing every data file you have currently downloaded and could be very time consuming.
To avoid this warning in the future, make sure that
/tmp/tmpeiy1ys_0/_downloaded_data.json
is not deleted between instantiations of this cache
warnings.warn(msg, MissingLocalManifestWarning)
ophys_session_table.csv: 100%|██████████| 247k/247k [00:00<00:00, 1.63MMB/s]
behavior_session_table.csv: 100%|██████████| 1.59M/1.59M [00:00<00:00, 7.10MMB/s]
ophys_experiment_table.csv: 100%|██████████| 657k/657k [00:00<00:00, 4.52MMB/s]
ophys_cells_table.csv: 100%|██████████| 4.28M/4.28M [00:00<00:00, 11.8MMB/s]
/home/runner/work/AllenSDK/AllenSDK/allensdk/brain_observatory/behavior/behavior_project_cache/behavior_project_cache.py:135: UpdatedStimulusPresentationTableWarning:
As of AllenSDK version 2.16.0, the latest Visual Behavior Ophys data has been significantly updated from previous releases. Specifically the user will need to update all processing of the stimulus_presentations tables. These tables now include multiple stimulus types delineated by the columns `stimulus_block` and `stimulus_block_name`.
The data that was available in previous releases are stored in the block name containing 'change_detection' and can be accessed in the pandas table by using:
`stimulus_presentations[stimulus_presentations.stimulus_block_name.str.contains('change_detection')]`
warnings.warn(
The data manifest is comprised of three types of tables:
behavior_session_tableophys_session_tableophys_experiment_table
The behavior_session_table contains metadata for every behavior session in the dataset. Some behavior sessions have 2-photon data associated with them, while others took place during training in the behavior facility. The different training stages that mice are progressed through are described by the session_type.
The ophys_session_table contains metadata for every 2-photon imaging (aka optical physiology, or ophys) session in the dataset, associated with a unique ophys_session_id. An ophys session is one continuous recording session under the microscope, and can contain different numbers of imaging planes (aka experiments) depending on which microscope was used. For Scientifica sessions, there will only be one experiment (aka imaging plane) per session. For Multiscope sessions, there can be up to eight imaging planes per session. Quality Control (QC) is performed on each individual imaging plane within a session, so each can fail QC independent of the others. This means that a Multiscope session may not have exactly eight experiments (imaging planes).
The ophys_experiment_table contains metadata for every ophys experiment in the dataset, which corresponds to a single imaging plane recorded in a single session, and associated with a unique ophys_experiment_id. A key part of our experimental design is targeting a given population of neurons, contained in one imaging plane, across multiple session_types (further described below) to examine the impact of varying sensory and behavioral conditions on single cell responses. The collection of all imaging sessions for a given imaging plane is referred to as an ophys container, associated with a unique ophys_container_id. Each ophys container may contain different numbers of sessions, depending on which experiments passed QC, and how many retakes occured (when a given session_type fails QC on the first try, an attempt is made to re-acquire the session_type on a different recording day - this is called a retake, also described further below).
To understand the difference between an ophys_experiment, an ophys_session, and an ophys_container, the following schematic can be helpful¶
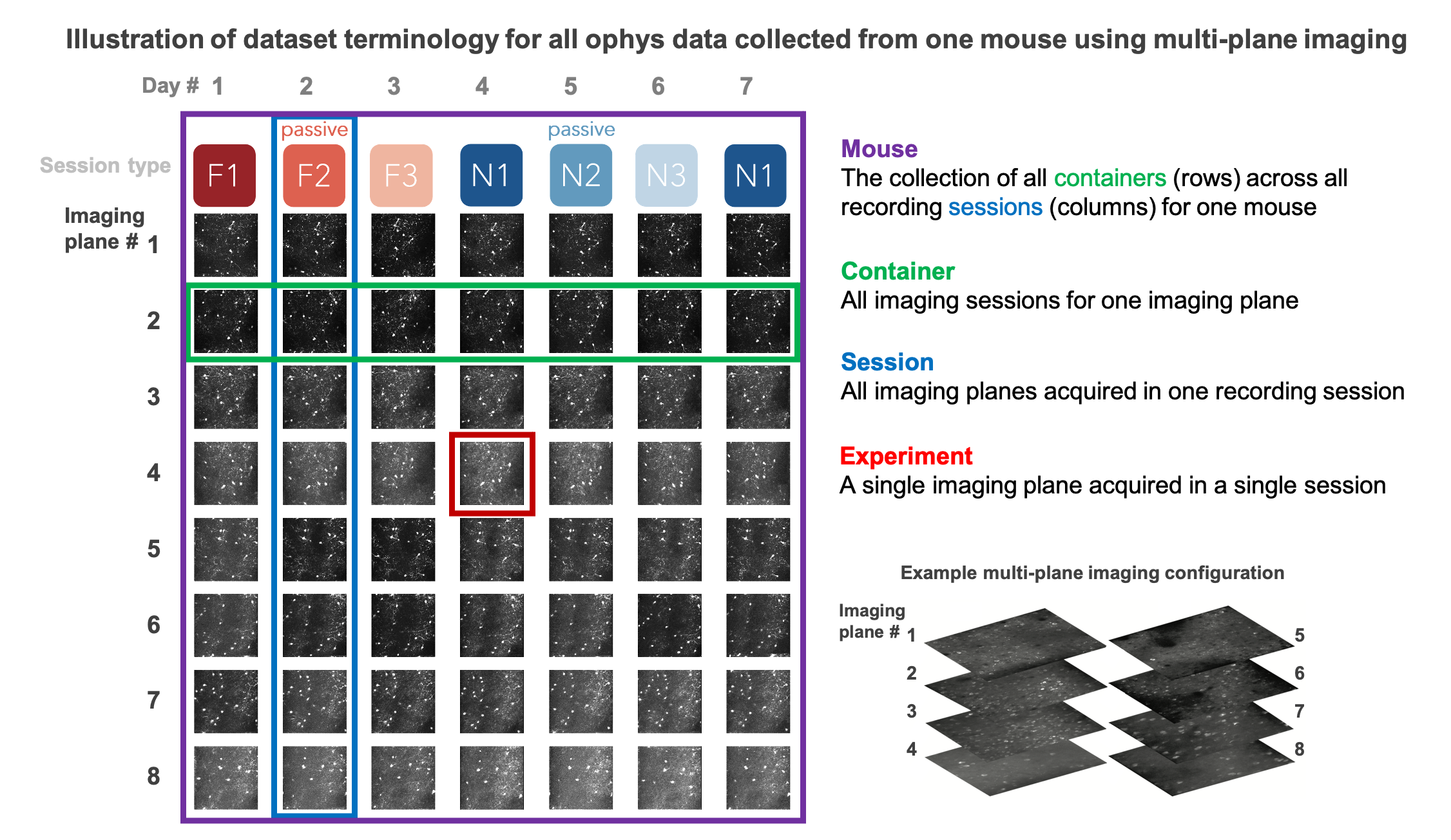
Note that this represents a multi-plane imaging dataset. For single-plane imaging, there will only be one plane, corresponding to one row of this diagram.
Lets go through each table and examine what metadata columns are available¶
Behavior Sessions Table¶
In this dataset, mice are trained on a visual change detection task. This task involves a continuous stream of stimuli, and mice learn to lick in response to a change in the stimulus identity to earn a water reward. There are different stages of training in this task, described below. The metadata for each behavior session in the dataset can be found in the behavior_sessions_table and can be used to identify behavior sessions you may want to analyze.
Load the behavior_sessions_table from the cache¶
behavior_sessions = cache.get_behavior_session_table()
print(f"Total number of behavior sessions: {len(behavior_sessions)}")
behavior_sessions.head()
Total number of behavior sessions: 4782
| ophys_session_id | ophys_container_id | mouse_id | indicator | full_genotype | driver_line | cre_line | reporter_line | sex | age_in_days | ... | num_targeted_structures | catch_trial_count | false_alarm_trial_count | miss_trial_count | correct_reject_trial_count | file_id | hit_trial_count | engaged_trial_count | go_trial_count | trial_count | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| behavior_session_id | |||||||||||||||||||||
| 870987812 | <NA> | NaN | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 80 | ... | <NA> | 0 | 0 | 0 | 0 | 1941 | 0 | 70 | 0 | 121 |
| 872953842 | <NA> | NaN | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 84 | ... | <NA> | 49 | 4 | 275 | 45 | 1942 | 18 | 27 | 293 | 564 |
| 873774125 | <NA> | NaN | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 85 | ... | <NA> | 41 | 3 | 300 | 38 | 1943 | 14 | 0 | 314 | 548 |
| 874790261 | <NA> | NaN | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 86 | ... | <NA> | 42 | 0 | 312 | 42 | 1944 | 14 | 0 | 326 | 492 |
| 875207834 | <NA> | NaN | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 87 | ... | <NA> | 44 | 3 | 300 | 41 | 1945 | 22 | 5 | 322 | 504 |
5 rows × 34 columns
What columns does the behavior_session table have and what values can they take?¶
behavior_sessions.columns
Index(['ophys_session_id', 'ophys_container_id', 'mouse_id', 'indicator',
'full_genotype', 'driver_line', 'cre_line', 'reporter_line', 'sex',
'age_in_days', 'imaging_plane_group_count', 'project_code',
'session_type', 'session_number', 'image_set', 'behavior_type',
'experience_level', 'prior_exposures_to_session_type',
'prior_exposures_to_image_set', 'prior_exposures_to_omissions',
'date_of_acquisition', 'equipment_name', 'num_depths_per_area',
'ophys_experiment_id', 'num_targeted_structures', 'catch_trial_count',
'false_alarm_trial_count', 'miss_trial_count',
'correct_reject_trial_count', 'file_id', 'hit_trial_count',
'engaged_trial_count', 'go_trial_count', 'trial_count'],
dtype='object')
behavior sessions can take place on different experimental systems¶
print('behavior data could be recorded on these experimental systems:\n')
print(np.sort(behavior_sessions.equipment_name.unique()))
behavior data could be recorded on these experimental systems: ['BEH.B' 'BEH.B-Box1' 'BEH.B-Box2' 'BEH.B-Box3' 'BEH.B-Box4' 'BEH.B-Box5' 'BEH.B-Box6' 'BEH.D' 'BEH.D-Box1' 'BEH.D-Box2' 'BEH.D-Box3' 'BEH.D-Box4' 'BEH.D-Box5' 'BEH.D-Box6' 'BEH.F' 'BEH.F-Box1' 'BEH.F-Box2' 'BEH.F-Box3' 'BEH.F-Box4' 'BEH.F-Box5' 'BEH.F-Box6' 'BEH.G' 'BEH.G-Box1' 'BEH.G-Box2' 'BEH.G-Box3' 'BEH.G-Box4' 'BEH.G-Box5' 'BEH.G-Box6' 'CAM2P.3' 'CAM2P.4' 'CAM2P.5' 'MESO.1']
equipment_name values starting with 'BEH' indicate behavioral training in the behavior facility, while values starting with 'CAM2P' or 'MESO' indicate behavior sessions that took place under a 2-photon microscope - either a Scientifica single plane imaging system ('CAMP2P.4', 'CAM2P.4', or 'CAM2P.5') or a modified Mesoscope system, also called Multiscope, for multi-plane imaging ('MESO.1').
Mouse specific metadata¶
The mouse_id is a 6-digit unique identifier for each experimental animal in the dataset
print('there are ', len(behavior_sessions.mouse_id.unique()), 'mice in the dataset')
there are 107 mice in the dataset
The transgenic line determines which neurons are labeled in a given mouse, and what they are labeled with¶
print('the different transgenic lines included in this dataset are:\n')
print(np.sort(behavior_sessions.full_genotype.unique()))
the different transgenic lines included in this dataset are: ['Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-GCaMP6f)/wt' 'Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai94(TITL-GCaMP6s)/wt' 'Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt' 'Vip-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt']
full_genotype refers to the full name of the transgenic mouse line, including all driver and reporter lines in the cross. driver_line and reporter_line have their own unique columns in the table. The first element of the full_genotype is the cre_line (which also has its own column in the table, and is a subset of driver_line). The cre_line determines which genetically identified neuron type will be labeled by the reporter_line.
print('the different cre lines used in this dataset are:\n')
print(np.sort(behavior_sessions.cre_line.unique()))
the different cre lines used in this dataset are: ['Slc17a7-IRES2-Cre' 'Sst-IRES-Cre' 'Vip-IRES-Cre']

In this dataset, we have 3 cre_lines, 'Slc17a7-IRES2-Cre', which labels excitatory neurons across all cortical layers, 'Sst-IRES-Cre' which labels somatostatin expressing inhibitory interneurons, and 'Vip-IRES-Cre', which labels vasoactive intestinal peptide expressing inhibitory interneurons. There are also 3 reporter_lines, 'Ai93(TITL-GCaMP6f)' which expresses the genetically encoded calcium indicator GCaMP6f (f is for 'fast', this reporter has fast offset kinetics, but is only moderately sensitive to calcium relative to other sensors) in cre labeled neurons, 'Ai94(TITL-GCaMP6s)' which expresses the indicator GCaMP6s (s is for 'slow', this reporter is very sensitive to calcium but has slow offset kinetics), and 'Ai148(TIT2L-GC6f-ICL-tTA2', which expresses GCaMP6f using a self-enhancing system to achieve higher expression than other reporter lines (which proved necessary to label inhibitory neurons specifically). The specific indicator expressed by each reporter_line also has its own column in the table.
print('the different reporter lines used in this dataset are:\n')
print(np.sort(behavior_sessions.reporter_line.unique()))
the different reporter lines used in this dataset are: ['Ai148(TIT2L-GC6f-ICL-tTA2)' 'Ai93(TITL-GCaMP6f)' 'Ai94(TITL-GCaMP6s)']
print('the different indicators used in this dataset are:\n')
print(np.sort(behavior_sessions.indicator.unique()))
the different indicators used in this dataset are: ['GCaMP6f' 'GCaMP6s']
- For more information about transgenic lines, see characterization data here: https://observatory.brain-map.org/visualcoding/transgenic
- for more information on GCaMP6, see this paper: https://www.nature.com/articles/nature12354
- For more information on reporter lines, see these papers: https://doi.org/10.1016/j.neuron.2015.02.022, https://www.sciencedirect.com/science/article/pii/S0092867418308031
how many mice per transgenic line?¶
behavior_sessions.groupby(['full_genotype', 'mouse_id']).count().reset_index().groupby('full_genotype').count()[['mouse_id']]
| mouse_id | |
|---|---|
| full_genotype | |
| Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-GCaMP6f)/wt | 47 |
| Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai94(TITL-GCaMP6s)/wt | 8 |
| Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | 24 |
| Vip-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | 28 |
Other mouse specific metadata includes sex and age_in_days
Session Type - a very important piece of information¶
The session_type for each behavior session indicates the behavioral training stage or 2-photon imaging conditions for that particular session. This determines what stimuli were shown and what task parameters were used.
print('the session_types available in this dataset are:\n')
print(np.sort(behavior_sessions.session_type[
~behavior_sessions.session_type.isna()].unique()))
the session_types available in this dataset are: ['OPHYS_0_images_A_habituation' 'OPHYS_0_images_B_habituation' 'OPHYS_0_images_G_habituation' 'OPHYS_1_images_A' 'OPHYS_1_images_B' 'OPHYS_1_images_G' 'OPHYS_2_images_A_passive' 'OPHYS_2_images_B_passive' 'OPHYS_2_images_G_passive' 'OPHYS_3_images_A' 'OPHYS_3_images_B' 'OPHYS_3_images_G' 'OPHYS_4_images_A' 'OPHYS_4_images_B' 'OPHYS_4_images_H' 'OPHYS_5_images_A_passive' 'OPHYS_5_images_B_passive' 'OPHYS_5_images_H_passive' 'OPHYS_6_images_A' 'OPHYS_6_images_B' 'OPHYS_6_images_H' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_A_10uL_reward' 'TRAINING_3_images_B_10uL_reward' 'TRAINING_3_images_G_10uL_reward' 'TRAINING_4_images_A_handoff_lapsed' 'TRAINING_4_images_A_handoff_ready' 'TRAINING_4_images_A_training' 'TRAINING_4_images_B_training' 'TRAINING_4_images_G_training' 'TRAINING_5_images_A_epilogue' 'TRAINING_5_images_A_handoff_lapsed' 'TRAINING_5_images_A_handoff_ready' 'TRAINING_5_images_B_epilogue' 'TRAINING_5_images_B_handoff_lapsed' 'TRAINING_5_images_B_handoff_ready' 'TRAINING_5_images_G_epilogue' 'TRAINING_5_images_G_handoff_lapsed' 'TRAINING_5_images_G_handoff_ready']
Mice are progressed through a series of training stages to shape their behavior prior to 2-photon imaging. Mice are automatically advanced between stages depending on their behavioral performance. For a detailed description of the change detection task and advancement criteria, please see the technical whitepaper: LINK
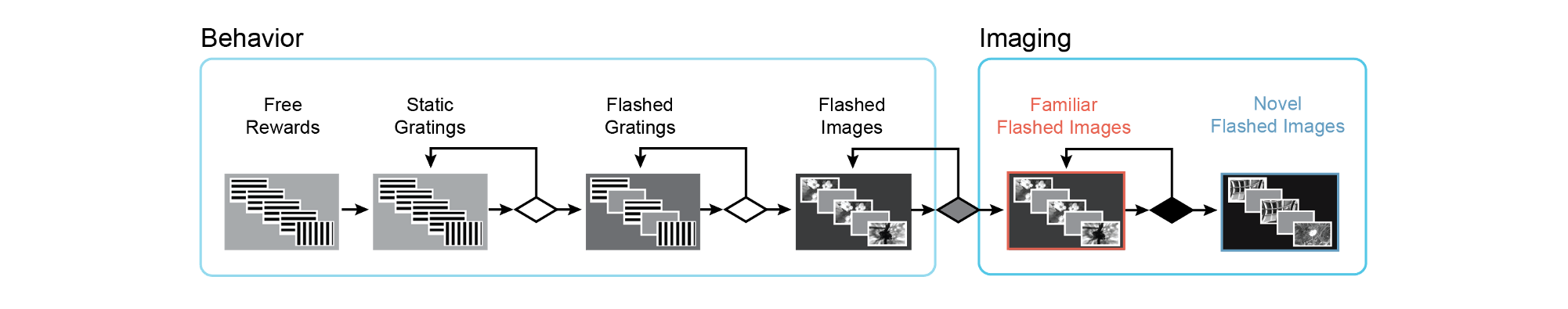
Training with the change detection task begins with simple static grating stimuli, changing between 0 and 90 degrees in orientation. On the very first day, mice are automatically given a water reward when the orientation of the stimulus changes (TRAINING_0_gratings_autorewards_15min). On subsequent days, mice must lick following the change in order to receive a water reward (TRAINING_1_gratings). In the next stage, stimuli are flashed, with a 500ms inter stimulus interal of mean luminance gray screen (TRAINING_2_gratings_flashed).
Once mice perform the task well with gratings, they are transitioned to natural image stimuli. Different groups of mice are trained with different sets of images, image set A or image set B (described further below). In the following description, we use X as a placeholder for image set A or B in the session_type name. Training with images begins with a 10ul water reward volume (TRAINING_3_images_X_10uL_reward), which is then decreased to 7ul once mice perform the task consistently with images (TRAINING_4_images_X_training). When mice have reached criterion to be transferred to the 2-photon imaging portion of the experiment, they are labeled as 'handoff_ready' (TRAINING_4_images_X_handoff_ready.) If behavior performance returns to below criterion level, they are labeled as 'handoff_lapsed'(TRAINING_4_images_X_handoff_lapsed).
# reminder about possible session types
print('the different session_types available in this dataset are:\n')
print(np.sort(behavior_sessions.session_type[
~behavior_sessions.session_type.isna()].unique()))
the different session_types available in this dataset are: ['OPHYS_0_images_A_habituation' 'OPHYS_0_images_B_habituation' 'OPHYS_0_images_G_habituation' 'OPHYS_1_images_A' 'OPHYS_1_images_B' 'OPHYS_1_images_G' 'OPHYS_2_images_A_passive' 'OPHYS_2_images_B_passive' 'OPHYS_2_images_G_passive' 'OPHYS_3_images_A' 'OPHYS_3_images_B' 'OPHYS_3_images_G' 'OPHYS_4_images_A' 'OPHYS_4_images_B' 'OPHYS_4_images_H' 'OPHYS_5_images_A_passive' 'OPHYS_5_images_B_passive' 'OPHYS_5_images_H_passive' 'OPHYS_6_images_A' 'OPHYS_6_images_B' 'OPHYS_6_images_H' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_A_10uL_reward' 'TRAINING_3_images_B_10uL_reward' 'TRAINING_3_images_G_10uL_reward' 'TRAINING_4_images_A_handoff_lapsed' 'TRAINING_4_images_A_handoff_ready' 'TRAINING_4_images_A_training' 'TRAINING_4_images_B_training' 'TRAINING_4_images_G_training' 'TRAINING_5_images_A_epilogue' 'TRAINING_5_images_A_handoff_lapsed' 'TRAINING_5_images_A_handoff_ready' 'TRAINING_5_images_B_epilogue' 'TRAINING_5_images_B_handoff_lapsed' 'TRAINING_5_images_B_handoff_ready' 'TRAINING_5_images_G_epilogue' 'TRAINING_5_images_G_handoff_lapsed' 'TRAINING_5_images_G_handoff_ready']
You will notice that some mice only go up to TRAINING_4, while others have the final training stage labeled TRAINING_5. This is due to a minor change made partway through data collection, where an epilogue stimulus was introduced during the final training stage prior to 2-photon imaging in order to habituate the mice to this stimulus, which is used during 2-photon imaging to aid in session to session registration. The epilogue is a 30 minute movie clip repeated 10 times, for a total of 5 minutes, and occurs at the end of the 60 minute behavioral session, followed by 5 minutes of blank gray screen. Training sessions with an epilogue movie include TRAINING_5_images_X_epilogue, TRAINING_5_images_X_handoff_ready , TRAINING_5_images_X_handoff_lapsed.
session_types during 2-photon imaging¶
When mice are transferred to the 2-photon rig for the imaging portion of the experiment, they first undergo 1-3 habituation sessions to get accustomed to the new experimental environment (OPHYS_0_images_X_habituation). During these sessions, mice perform the task under the microscope, but no experimental data is recorded.

During the 2-photon imaging portion of the experiment, mice perform the task with the same set of images they saw during training (either image set A or B), as well as an additional novel set of images (whichever of A or B that they did not see during training). This allows evaluation of the impact of different sensory contexts on neural activity - familiarity versus novelty. Sessions with familiar images include those starting with OPHYS_0, OPHYS_1, OPHYS_2, and OPHYS_3. Sessions with novel images include those starting with OPHYS_4, OPHYS_5, and OPHYS_6.
Interleaved between active behavior sessions are passive viewing sessions where mice are given their daily water ahead of the sesssion (and are thus satiated) and view the stimulus with the lick spout retracted so they are unable to earn water rewards. This allows comparison of neural activity in response to stimuli under different behavioral context - active task engagement and passive viewing without reward. Passive sessions include OPHYS_2_images_A_passive (passive session with familiar images), and OPHYS_5_images_A_passive (passive session with novel images).
The final session during the 2-photon imaging phase is OPHYS_7_receptive_field_mapping, however 2-photon data is not available for these sessions in this data release (but will be made available in a subsequent release).
Dataset variants - different mice were subject to different experimental conditions¶
As hinted to above, some mice were trained with image set A, and others with image set B. Including these two groups of mice, with swapped stimulus conditions, was included in the dataset as a control for the effects of novelty, to ensure that any observed changes were truly due to lack of familiarity with the novel image set, rather than a result of specific features of the image set that was used. In addition, some mice were imaged on the Scientifica single plane imaging systems, and other mice were imaged on Multiscope for multi-plane imaging. These distinct groups of mice are referred to as dataset variants and can be identified using the project_code column
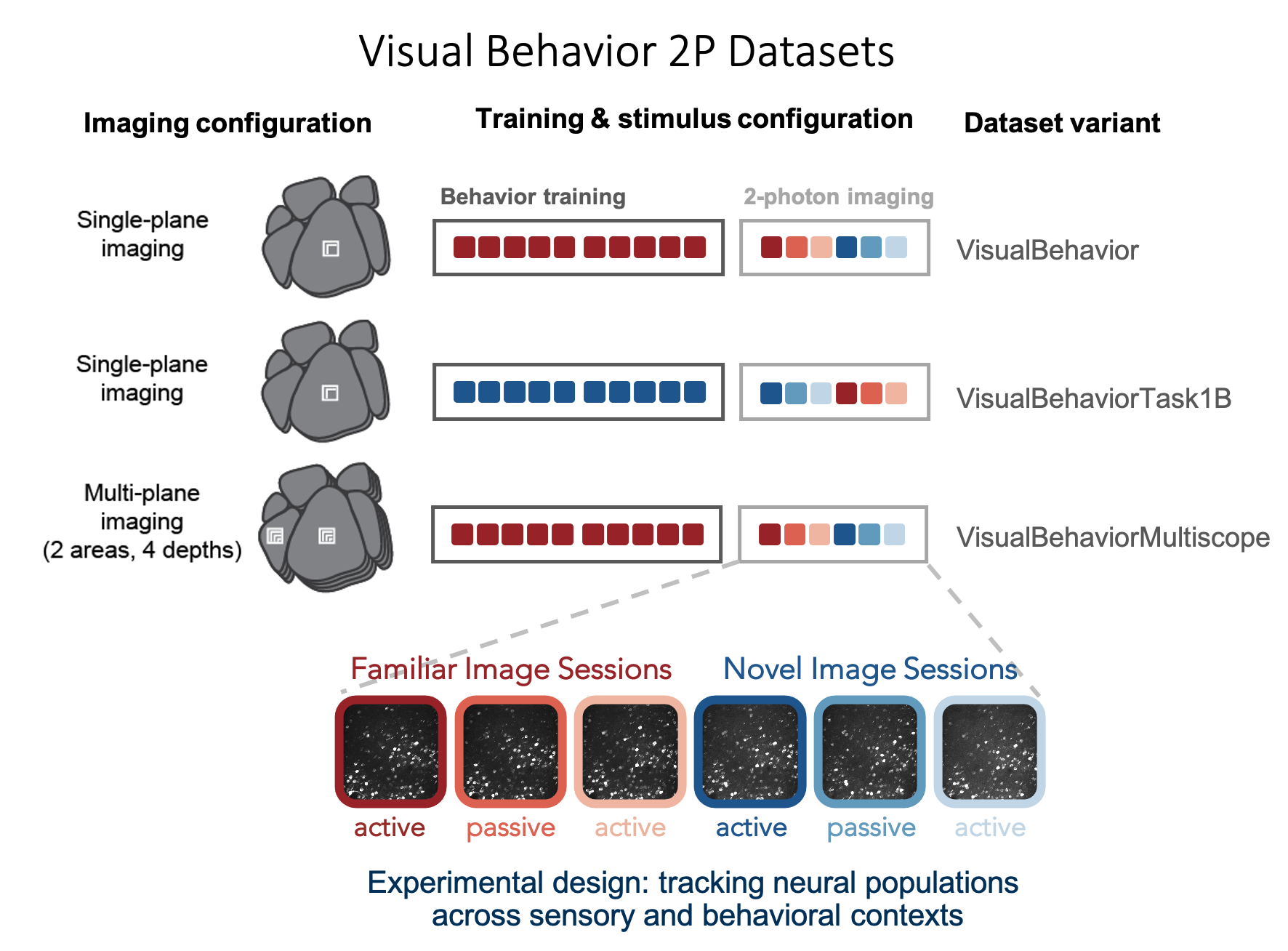
Project_code is only defined for ophys sessions, for technical reasons, so let's fill in the gaps so that all mice have a project_code
# get a table of the project code for each mouse
project_code_lookup = behavior_sessions[behavior_sessions.project_code.isnull()==False].reset_index().drop_duplicates('mouse_id')[['mouse_id','project_code']]
project_code_lookup
| mouse_id | project_code | |
|---|---|---|
| 0 | 457841 | VisualBehaviorMultiscope |
| 83 | 476970 | VisualBehaviorTask1B |
| 159 | 412036 | VisualBehavior |
| 206 | 429952 | VisualBehavior |
| 239 | 449441 | VisualBehavior |
| ... | ... | ... |
| 4494 | 467302 | VisualBehavior |
| 4551 | 489065 | VisualBehaviorMultiscope4areasx2d |
| 4612 | 403491 | VisualBehavior |
| 4683 | 523927 | VisualBehavior |
| 4753 | 489066 | VisualBehaviorTask1B |
107 rows × 2 columns
behavior_sessions = behavior_sessions.merge(project_code_lookup, on='mouse_id',
how='left', suffixes=('_session', '_mouse'))
behavior_sessions = behavior_sessions.drop(columns='project_code_session')
behavior_sessions = behavior_sessions.rename(columns={'project_code_mouse': 'project_code'})
What project_codes are available? What session_types belong to each?¶
behavior_sessions.project_code.unique()
array(['VisualBehaviorMultiscope', 'VisualBehaviorTask1B',
'VisualBehavior', 'VisualBehaviorMultiscope4areasx2d'],
dtype=object)
for project_code in behavior_sessions.project_code.unique():
project_sessions = behavior_sessions[behavior_sessions.project_code==project_code]
print('\n project_code:', project_code)
print('\n has these session types:\n', np.sort(
project_sessions.session_type[~project_sessions.session_type.isna()].unique()))
print('\n')
project_code: VisualBehaviorMultiscope has these session types: ['OPHYS_0_images_A_habituation' 'OPHYS_1_images_A' 'OPHYS_2_images_A_passive' 'OPHYS_3_images_A' 'OPHYS_3_images_B' 'OPHYS_4_images_B' 'OPHYS_5_images_A_passive' 'OPHYS_5_images_B_passive' 'OPHYS_6_images_A' 'OPHYS_6_images_B' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_A_10uL_reward' 'TRAINING_4_images_A_training' 'TRAINING_5_images_A_epilogue' 'TRAINING_5_images_A_handoff_lapsed' 'TRAINING_5_images_A_handoff_ready'] project_code: VisualBehaviorTask1B has these session types: ['OPHYS_0_images_A_habituation' 'OPHYS_0_images_B_habituation' 'OPHYS_1_images_B' 'OPHYS_2_images_B_passive' 'OPHYS_3_images_B' 'OPHYS_4_images_A' 'OPHYS_4_images_B' 'OPHYS_5_images_A_passive' 'OPHYS_6_images_A' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_B_10uL_reward' 'TRAINING_4_images_B_training' 'TRAINING_5_images_B_epilogue' 'TRAINING_5_images_B_handoff_lapsed' 'TRAINING_5_images_B_handoff_ready'] project_code: VisualBehavior has these session types: ['OPHYS_0_images_A_habituation' 'OPHYS_0_images_B_habituation' 'OPHYS_1_images_A' 'OPHYS_1_images_B' 'OPHYS_2_images_A_passive' 'OPHYS_2_images_B_passive' 'OPHYS_3_images_A' 'OPHYS_3_images_B' 'OPHYS_4_images_A' 'OPHYS_4_images_B' 'OPHYS_5_images_A_passive' 'OPHYS_5_images_B_passive' 'OPHYS_6_images_A' 'OPHYS_6_images_B' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_A_10uL_reward' 'TRAINING_3_images_B_10uL_reward' 'TRAINING_4_images_A_handoff_lapsed' 'TRAINING_4_images_A_handoff_ready' 'TRAINING_4_images_A_training' 'TRAINING_4_images_B_training' 'TRAINING_5_images_A_epilogue' 'TRAINING_5_images_A_handoff_lapsed' 'TRAINING_5_images_A_handoff_ready' 'TRAINING_5_images_B_epilogue' 'TRAINING_5_images_B_handoff_ready'] project_code: VisualBehaviorMultiscope4areasx2d has these session types: ['OPHYS_0_images_G_habituation' 'OPHYS_1_images_G' 'OPHYS_2_images_G_passive' 'OPHYS_3_images_G' 'OPHYS_4_images_H' 'OPHYS_5_images_H_passive' 'OPHYS_6_images_H' 'TRAINING_0_gratings_autorewards_15min' 'TRAINING_1_gratings' 'TRAINING_2_gratings_flashed' 'TRAINING_3_images_G_10uL_reward' 'TRAINING_4_images_G_training' 'TRAINING_5_images_G_epilogue' 'TRAINING_5_images_G_handoff_lapsed' 'TRAINING_5_images_G_handoff_ready']
Notice that for project_codes VisualBehavior and VisualBehaviorMultiscope, mice are trained on image set A, while for VisualBehaviorTask1B, mice are trained on image set B
Ophys Sessions Table¶
The ophys_session_table includes all of the metadata columns available in the behavior_session_table, as well as additional information specific to 2-photon imaging, namely the list of ophys_experiment_ids and ophys_container_ids associated with each ophys_session_id.
ophys_sessions = cache.get_ophys_session_table()
print(f"Total number of ophys sessions: {len(ophys_sessions)}\n")
print(ophys_sessions.columns)
ophys_sessions.head()
Total number of ophys sessions: 703
Index(['behavior_session_id', 'ophys_container_id', 'mouse_id', 'indicator',
'full_genotype', 'driver_line', 'cre_line', 'reporter_line', 'sex',
'age_in_days', 'imaging_plane_group_count', 'project_code',
'session_type', 'session_number', 'image_set', 'behavior_type',
'experience_level', 'prior_exposures_to_session_type',
'prior_exposures_to_image_set', 'prior_exposures_to_omissions',
'date_of_acquisition', 'equipment_name', 'num_depths_per_area',
'ophys_experiment_id', 'num_targeted_structures'],
dtype='object')
| behavior_session_id | ophys_container_id | mouse_id | indicator | full_genotype | driver_line | cre_line | reporter_line | sex | age_in_days | ... | behavior_type | experience_level | prior_exposures_to_session_type | prior_exposures_to_image_set | prior_exposures_to_omissions | date_of_acquisition | equipment_name | num_depths_per_area | ophys_experiment_id | num_targeted_structures | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ophys_session_id | |||||||||||||||||||||
| 951410079 | 951520319 | [1018028339, 1018028342, 1018028345, 101802835... | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 206 | ... | active_behavior | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 4 | [951980471, 951980473, 951980475, 951980479, 9... | 2 |
| 952430817 | 952554548 | [1018028339, 1018028345, 1018028354, 1018028357] | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 209 | ... | passive_viewing | Familiar | 0 | 66 | 1 | 2019-09-23 08:45:38.490000+00:00 | MESO.1 | 4 | [953659743, 953659745, 953659749, 953659752] | 2 |
| 954954402 | 953982960 | [1018028339, 1018028342, 1018028345, 101802835... | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 210 | ... | active_behavior | Familiar | 0 | 67 | 2 | 2019-09-24 09:01:31.582000+00:00 | MESO.1 | 4 | [958527464, 958527471, 958527474, 958527479, 9... | 2 |
| 955775716 | 956010809 | [1018028339, 1018028342, 1018028345] | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 212 | ... | active_behavior | Familiar | 1 | 68 | 3 | 2019-09-26 09:22:21.772000+00:00 | MESO.1 | 4 | [956941841, 956941844, 956941846] | 2 |
| 957020350 | 957032492 | [1018028339, 1018028342, 1018028345, 101802835... | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | 213 | ... | active_behavior | Novel 1 | 0 | 0 | 4 | 2019-09-27 08:58:37.005000+00:00 | MESO.1 | 4 | [957759562, 957759564, 957759566, 957759570, 9... | 2 |
5 rows × 25 columns
# what do the ophys_experiment_id and ophys_container_id columns look like?
# are there always the same number of experiments and containers in different sessions?
# does the number of experiments and containers depend on the microscope used?
ophys_sessions[['ophys_experiment_id', 'ophys_container_id', 'equipment_name']][:15]
| ophys_experiment_id | ophys_container_id | equipment_name | |
|---|---|---|---|
| ophys_session_id | |||
| 951410079 | [951980471, 951980473, 951980475, 951980479, 9... | [1018028339, 1018028342, 1018028345, 101802835... | MESO.1 |
| 952430817 | [953659743, 953659745, 953659749, 953659752] | [1018028339, 1018028345, 1018028354, 1018028357] | MESO.1 |
| 954954402 | [958527464, 958527471, 958527474, 958527479, 9... | [1018028339, 1018028342, 1018028345, 101802835... | MESO.1 |
| 955775716 | [956941841, 956941844, 956941846] | [1018028339, 1018028342, 1018028345] | MESO.1 |
| 957020350 | [957759562, 957759564, 957759566, 957759570, 9... | [1018028339, 1018028342, 1018028345, 101802835... | MESO.1 |
| 958105827 | [958741219, 958741222, 958741230, 958741232, 9... | [1018028342, 1018028345, 1018028351, 101802835... | MESO.1 |
| 958772311 | [959388788, 959388790, 959388792, 959388796, 9... | [1018028339, 1018028342, 1018028345, 101802835... | MESO.1 |
| 959458018 | [960410023, 960410026, 960410028, 960410038, 9... | [1018028339, 1018028342, 1018028345, 101802835... | MESO.1 |
| 993727065 | [993862620] | [991913453] | CAM2P.4 |
| 993984066 | [994053903] | [991913453] | CAM2P.4 |
| 994199725 | [994278281] | [991913453] | CAM2P.4 |
| 994731371 | [994790561] | [991913453] | CAM2P.4 |
| 994883056 | [994955425] | [991913453] | CAM2P.4 |
| 995115654 | [995280513] | [991913453] | CAM2P.4 |
| 995359867 | [995439942] | [991913453] | CAM2P.4 |
Session order¶
The ophys_session_table only includes sessions that pass ophys QC¶
(but the behavior_session_table includes all the sessions)¶
The ophys_session_table only includes sessions with 2-photon imaging data that passed our QC criteria. Importantly, sessions that took place during 2-photon imaging, but did NOT pass QC, can be found in the behavior_session_table, as it includes the full training history for every mouse. In the behavior_session_table, only sessions with passing ophys data will have an ophys_session_id. We can use this to identify ophys sessions that didnt pass QC, but still have behavior data.
Let's look at all the behavior sessions that took place on a 2-photon rig for one mouse, in order of acquisition date¶
# pick a mouse
mouse_id = 445002
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='CAM2P.3')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['session_type', 'date_of_acquisition', 'ophys_session_id']]
| session_type | date_of_acquisition | ophys_session_id |
|---|
Notice that only a subset of all OPHYS sessions have an ophys_session_id - these are the sessions that passed QC. Sessions with NaN as the ophys_session_id either do not have 2P data recorded (as in habituation sessions), or failed QC and were retaken on a subsequent day, such as OPHYS_5_images_B_passive in this case
print('there are', len(mouse_ophys_sessions), 'ophys sessions in the behavior_session_table for this mouse')
print('this includes ophys sessions that failed QC for ophys, but still have behavior data')
there are 0 ophys sessions in the behavior_session_table for this mouse this includes ophys sessions that failed QC for ophys, but still have behavior data
What is available in the ophys_session_table for this mouse?¶
print('there are', len(ophys_sessions[ophys_sessions.mouse_id==mouse_id]), 'sessions in the ophys_session_table for this mouse')
print('these are the sessions with valid ophys data')
there are 0 sessions in the ophys_session_table for this mouse these are the sessions with valid ophys data
ophys_sessions[ophys_sessions.mouse_id==mouse_id][['date_of_acquisition', 'session_type']]
| date_of_acquisition | session_type | |
|---|---|---|
| ophys_session_id |
Due to QC failures and retakes, session types in the ophys_session_table do not always occur in sequential order¶
The schematic above depicts ophys sessions OPHYS1-6 in a specific order, however this order is rarely perfectly maintained due to QC failures. The example above shows OPHYS_1-4 in the correct order, but then OPHYS_5 comes after OPHYS_6 because the first attempt at OPHYS_5 failed (as we can see from the behavior_sessions for this mouse), and had to be retaken after OPHYS_6.
Let's look at the session order for a different mouse, imaged on the Multiscope¶
# pick a mouse
mouse_id = 453911
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='MESO.1')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['date_of_acquisition', 'session_type', 'ophys_session_id']]
| date_of_acquisition | session_type | ophys_session_id |
|---|
Looks like lots of retakes for this one (where ophys_session_id = NaN). Also note that there are multiple retakes for some session_types. This can happen for mice imaged on Multiscope, because retakes can be triggered by QC failure of any one of the 8 imaging planes in the session.
Let's look at how failures and retakes affects the session order in the ophys_sessions table for this mouse¶
ophys_sessions[ophys_sessions.mouse_id==mouse_id][['date_of_acquisition', 'session_type']]
| date_of_acquisition | session_type | |
|---|---|---|
| ophys_session_id |
It looks like the first set of sessions are taken in sequential order, but after that there are a few retakes of some of the session_types. This suggests that some of the imaging planes for this Multiscope mouse passed QC on the first time around, but retakes were needed to get passing ophys data for other imaging planes.
But they're not always out of order, sometimes things go perfectly!¶
# pick a mouse
mouse_id = 438912
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='MESO.1')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['date_of_acquisition', 'session_type', 'ophys_session_id']]
| date_of_acquisition | session_type | ophys_session_id |
|---|
Well, nearly perfectly, OPHYS_5 came after OPHYS_6
Prior Exposures¶
Because the session types can be out of order due to retakes, and because of some of our other experimental design decisions, it is helpful to know some information about the history of the mouse relative to a given session. To serve this purpose, we have included metadata describing the prior_exposures_to_image_set, prior_exposures_to_session_type, and prior_exposures_to_omissions as columns in all the manifest data tables.
prior_exposures_to_image_set¶
A key aspect of our experimental design is the inclusion of novel stimuli during the imaging phase of the experiment. However, after the very first session with these novel images, they actually start to become more and more familiar. So, it is important to know whether a given session is truly the first exposure to that image set. In addition, it is useful to know whether subsequent sessions are the second, third, fourth, etc. exposure to that image set, for analysis of changes in activity with experience following novelty. The prior_exposures_to_image_set column describes the number of sessions that a given mouse has observed the stimulus set that was shown in that session, prior to the start of that session. For the very first exposure to a novel image set, the value of prior_exposures_to_image_set will be 0.
Let's look at the prior_exposures_to_image_set column for one of the mice we looked at above, first in the behavior_session_table, which contains all sessions the mouse experienced, then in the ophys_session_table, which only includes sessions that passed ophys QC¶
mouse_id = 445002
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='CAM2P.3')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['session_type', 'date_of_acquisition', 'ophys_session_id', 'prior_exposures_to_image_set']]
| session_type | date_of_acquisition | ophys_session_id | prior_exposures_to_image_set |
|---|
Note that prior_exposures_to_image_set is a high number for OPHYS_0-3, because that is the image set the mouse was trained on, and that it re-sets to zero for the first exposure to the novel image set in OPHYS_4
Let's double check the full training history for this mouse¶
mouse_id = 445002
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['session_type', 'date_of_acquisition', 'ophys_session_id', 'prior_exposures_to_image_set']]
| session_type | date_of_acquisition | ophys_session_id | prior_exposures_to_image_set |
|---|
Knowing the prior exposures number is especially important for the ophys_session_table, because the sessions that failed ophys QC are not visible there, so it is difficult to know whether a given session was the first of its type or a retake.
ophys_sessions[ophys_sessions.mouse_id==mouse_id][['date_of_acquisition', 'session_type', 'prior_exposures_to_image_set']]
| date_of_acquisition | session_type | prior_exposures_to_image_set | |
|---|---|---|---|
| ophys_session_id |
prior_exposures_to_session_type¶
In some cases, you may want to know how many times a given session_type was seen by the mouse. For example, to know whether a passive viewing session was the very first time the mouse experienced a passive session with no lick spout, as there may be a difference in expectation of reward between the first passive session and a later one where the mouse has become accustomed to sometimes having the lick spout removed.
compare prior_exposures_to_session_type in the behavior_session_table with the ophys_session_table for a given mouse¶
# pick a mouse
mouse_id = 456915
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='MESO.1')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['date_of_acquisition', 'session_type', 'ophys_session_id', 'prior_exposures_to_session_type']]
| date_of_acquisition | session_type | ophys_session_id | prior_exposures_to_session_type |
|---|
ophys_sessions[ophys_sessions.mouse_id==mouse_id][['date_of_acquisition', 'session_type', 'prior_exposures_to_session_type']]
| date_of_acquisition | session_type | prior_exposures_to_session_type | |
|---|---|---|---|
| ophys_session_id |
Without the prior_exposures_to_session_type column in the ophys_session_table, it would be difficult to know that OPHYS_2_images_A_passive was actually the second time (1 prior exposure) that the mouse had experienced a passive vieweing session
prior_exposures_to_omissions¶
Another unique aspect of the experimental design of this dataset is the inclusion of stimulus omissions in the 2-photon portion of the experiment. During behavioral training, mice experience a highly regular cadence of himage presentations, with 250ms per stimulus presentation, with a 500ms gray screen in between. During imaging sessions, stimulus presentations (other than the change and pre-change images) are omitted with a 5% probability, resulting in some inter stimlus intervals appearing as an extended gray screen period. This allows exploration of potential effects of temporal expectation on neural activity.
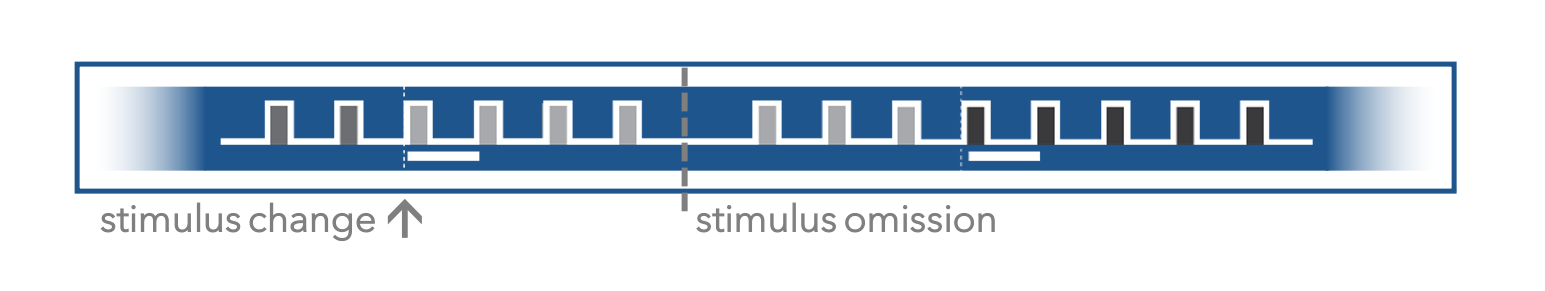
Let's look at prior_exposures_to_omissions in a few mice¶
np.sort(behavior_sessions[behavior_sessions.equipment_name=='CAM2P.4'].mouse_id.unique())
array(['412036', '412366', '423606', '425493', '425496', '429952',
'431252', '436662', '442709', '447663', '448900', '453913',
'456564', '462468', '467953', '468866', '476970', '479458',
'480753', '489066', '492395', '523927', '524274', '533161',
'533162', '538219', '539518', '544261', '549976'], dtype=object)
# pick a mouse
mouse_id = 436662
# get behavior sessions that took place on the microscope
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)&
(behavior_sessions.equipment_name=='CAM2P.4')]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['date_of_acquisition', 'session_type', 'ophys_session_id', 'equipment_name', 'prior_exposures_to_omissions']]
| date_of_acquisition | session_type | ophys_session_id | equipment_name | prior_exposures_to_omissions |
|---|
In this case (and in most cases), omissions do not occur until the first true imaging session on the 2-photon rig, OPHYS_1, i.e. they are not included in habituation sessions. However, in a small number of mice from the beginning of our data collection process, omissions did occur in habituation sessions (but never during training). This is something to be careful of if you are looking at something like the change in omission related activity with experience.
Here is a mouse that saw omissions during habituation sessions. Also note that the first two habituation sessions took place on different microscopes (this is extremely rare, every effort is made to image a given mouse on the same 2-photon rig during its entire lifetime).
# pick a mouse
mouse_id = 423606
# get behavior sessions - include training as well
mouse_ophys_sessions = behavior_sessions[(behavior_sessions.mouse_id==mouse_id)]
# only look at the relevant columns
mouse_ophys_sessions.sort_values(by='date_of_acquisition')[['date_of_acquisition', 'session_type', 'ophys_session_id', 'equipment_name', 'prior_exposures_to_omissions']]
| date_of_acquisition | session_type | ophys_session_id | equipment_name | prior_exposures_to_omissions |
|---|
Here is how to identify all mice that saw omissions during habituation sessions¶
# get all behavior sessions that were habituation sessions (image set A or B)
# where the prior exposures to omissions was not zero
habituation_with_omission = behavior_sessions[((behavior_sessions.session_type=='OPHYS_0_images_A_habituation')|
(behavior_sessions.session_type=='OPHYS_0_images_B_habituation'))&
(behavior_sessions.prior_exposures_to_omissions>0)]
mice_with_omission_during_habituation = habituation_with_omission.mouse_id.unique()
print(len(mice_with_omission_during_habituation), ' mice had omissions during habituation')
14 mice had omissions during habituation
Ophys Experiment Table¶
The ophys_experiment_table contains all ophys data that passes QC, organized according to individual imaging planes in individual sessions, each associated with an ophys_experiment_id. The ophys_experiment_table contains all the columns in ophys_session_table, plus a few additional ones specific to individual imaging planes, namely imaging_depth and targeted_structure.
ophys_experiments = cache.get_ophys_experiment_table()
print(f"Total number of ophys experiments: {len(ophys_experiments)}\n")
ophys_experiments.head()
Total number of ophys experiments: 1936
| behavior_session_id | ophys_session_id | ophys_container_id | mouse_id | indicator | full_genotype | driver_line | cre_line | reporter_line | sex | ... | passive | experience_level | prior_exposures_to_session_type | prior_exposures_to_image_set | prior_exposures_to_omissions | date_of_acquisition | equipment_name | published_at | isi_experiment_id | file_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ophys_experiment_id | |||||||||||||||||||||
| 951980471 | 951520319 | 951410079 | 1018028342 | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 2021-03-25 | 848974280 | 0 |
| 951980473 | 951520319 | 951410079 | 1018028345 | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 2021-03-25 | 848974280 | 1 |
| 951980475 | 951520319 | 951410079 | 1018028339 | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 2021-03-25 | 848974280 | 2 |
| 951980479 | 951520319 | 951410079 | 1018028354 | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 2021-03-25 | 848974280 | 3 |
| 951980481 | 951520319 | 951410079 | 1018028357 | 457841 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 65 | 0 | 2019-09-20 09:59:38.837000+00:00 | MESO.1 | 2021-03-25 | 848974280 | 4 |
5 rows × 30 columns
Compare the columns of ophys_sessions_table with ophys_experiments_table¶
ophys_sessions.columns
Index(['behavior_session_id', 'ophys_container_id', 'mouse_id', 'indicator',
'full_genotype', 'driver_line', 'cre_line', 'reporter_line', 'sex',
'age_in_days', 'imaging_plane_group_count', 'project_code',
'session_type', 'session_number', 'image_set', 'behavior_type',
'experience_level', 'prior_exposures_to_session_type',
'prior_exposures_to_image_set', 'prior_exposures_to_omissions',
'date_of_acquisition', 'equipment_name', 'num_depths_per_area',
'ophys_experiment_id', 'num_targeted_structures'],
dtype='object')
ophys_experiments.columns
Index(['behavior_session_id', 'ophys_session_id', 'ophys_container_id',
'mouse_id', 'indicator', 'full_genotype', 'driver_line', 'cre_line',
'reporter_line', 'sex', 'age_in_days', 'imaging_depth',
'targeted_structure', 'targeted_imaging_depth', 'imaging_plane_group',
'project_code', 'session_type', 'session_number', 'image_set',
'behavior_type', 'passive', 'experience_level',
'prior_exposures_to_session_type', 'prior_exposures_to_image_set',
'prior_exposures_to_omissions', 'date_of_acquisition', 'equipment_name',
'published_at', 'isi_experiment_id', 'file_id'],
dtype='object')
What imaging_depths and targeted_structures are available? Are they different depending on project_code?¶
# loop through project codes and print the available imaging_depths and targeted_structures
for project_code in ophys_experiments.project_code.unique():
project_experiments = ophys_experiments[ophys_experiments.project_code==project_code]
print('\nimaging_depths available for', project_code, 'include: ', project_experiments.imaging_depth.unique())
print('\ntargeted_structures available for', project_code, 'include: ', project_experiments.targeted_structure.unique())
print('\n')
imaging_depths available for VisualBehaviorMultiscope include: [150 225 75 300 169 263 73 359 271 85 267 361 167 275 79 363 177 269 365 175 283 367 165 373 81 371 369 171 259 77 279 83 377 274 152 71 146 227 158 144 276 160 87 142 217 69 51 140 219 215 156 211 162 148 223 166 231 174 282 93 213 221 229 306 320 298 232 218 179 285 270 272 145 251 273 154 308 302 278 312 383 161 381 265 261 230 290 228 61 284 233 304 292 65 289 235 310 375 280 281 187 91 197 295 183 355 277 181 185 291 95 173 67 321 149 287 322 220 226 222 315 147 294 286 132 316 296 350 348 379 340 345] targeted_structures available for VisualBehaviorMultiscope include: ['VISp' 'VISl'] imaging_depths available for VisualBehaviorTask1B include: [275 175 375] targeted_structures available for VisualBehaviorTask1B include: ['VISp'] imaging_depths available for VisualBehavior include: [175 375 275 290] targeted_structures available for VisualBehavior include: ['VISp'] imaging_depths available for VisualBehaviorMultiscope4areasx2d include: [187 163 263 179 265 167 267 183 171 255 181 169 271 175 273 277 275 269 193 185 189 283 177 165 173 159 279 281 150 260 240 160 205 155 210 145 125 270 300 182 174 257 285 170 245 295 195 250 156 251 297 249 190 305 176 244 200 310 107 153 227 123 103 157 111 161 139 225 109 110 137 221 101 135 219 121 151 105 261 213 217 143 211 133 241 117 147 223 287 308 178 180 290 286 280 268 141 253 289 201 254 235 230 128 224 132 222 233 229 130 124 236 134 191] targeted_structures available for VisualBehaviorMultiscope4areasx2d include: ['VISp' 'VISl' 'VISal' 'VISam']
ophys_experiment_table is useful for identifying ophys_containers to analyze¶
Compare the ophys_container_id column of the ophys_experiment_table with the ophys_session_table. In ophys_session_table, each ophys_session_id is associated with one or more imaging planes (ophys_experiment_ids), while in the ophys_experiment_table, you can evaluate each of those imaging planes indepdently. This is particularly helpful for identifying ophys_containers that you want to analyze - the set of all imaging sessions for a given imaging plane.
The ophys_experient_table has all the same columns as ophys_session_table, just reorgnized by ophys_experiment_id
print(ophys_experiments.columns)
Index(['behavior_session_id', 'ophys_session_id', 'ophys_container_id',
'mouse_id', 'indicator', 'full_genotype', 'driver_line', 'cre_line',
'reporter_line', 'sex', 'age_in_days', 'imaging_depth',
'targeted_structure', 'targeted_imaging_depth', 'imaging_plane_group',
'project_code', 'session_type', 'session_number', 'image_set',
'behavior_type', 'passive', 'experience_level',
'prior_exposures_to_session_type', 'prior_exposures_to_image_set',
'prior_exposures_to_omissions', 'date_of_acquisition', 'equipment_name',
'published_at', 'isi_experiment_id', 'file_id'],
dtype='object')
This means that each ophys_experiment_id has a single ophys_container_id.
Let's pick an ophys_container_id and see what ophys_experiments it contains?¶
ophys_container_id = ophys_experiments.ophys_container_id.unique()[50]
container_experiments = ophys_experiments[ophys_experiments.ophys_container_id==ophys_container_id]
container_experiments
| behavior_session_id | ophys_session_id | ophys_container_id | mouse_id | indicator | full_genotype | driver_line | cre_line | reporter_line | sex | ... | passive | experience_level | prior_exposures_to_session_type | prior_exposures_to_image_set | prior_exposures_to_omissions | date_of_acquisition | equipment_name | published_at | isi_experiment_id | file_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ophys_experiment_id | |||||||||||||||||||||
| 975608418 | 975239345 | 975050422 | 1018027765 | 484408 | GCaMP6f | Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-G... | [Slc17a7-IRES2-Cre, Camk2a-tTA] | Slc17a7-IRES2-Cre | Ai93(TITL-GCaMP6f) | M | ... | False | Familiar | 0 | 9 | 1 | 2019-11-01 09:25:23.468000+00:00 | MESO.1 | 2021-08-12 | 942701302 | 293 |
| 976300323 | 976039179 | 975929719 | 1018027765 | 484408 | GCaMP6f | Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-G... | [Slc17a7-IRES2-Cre, Camk2a-tTA] | Slc17a7-IRES2-Cre | Ai93(TITL-GCaMP6f) | M | ... | True | Familiar | 0 | 10 | 2 | 2019-11-04 09:21:00.916000+00:00 | MESO.1 | 2021-08-12 | 942701302 | 301 |
| 977220493 | 976964099 | 976865980 | 1018027765 | 484408 | GCaMP6f | Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-G... | [Slc17a7-IRES2-Cre, Camk2a-tTA] | Slc17a7-IRES2-Cre | Ai93(TITL-GCaMP6f) | M | ... | False | Familiar | 0 | 11 | 3 | 2019-11-05 13:35:33.124000+00:00 | MESO.1 | 2021-08-12 | 942701302 | 309 |
| 977975802 | 977451727 | 977288921 | 1018027765 | 484408 | GCaMP6f | Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-G... | [Slc17a7-IRES2-Cre, Camk2a-tTA] | Slc17a7-IRES2-Cre | Ai93(TITL-GCaMP6f) | M | ... | False | Novel 1 | 0 | 0 | 4 | 2019-11-06 09:02:04.020000+00:00 | MESO.1 | 2021-08-12 | 942701302 | 317 |
| 978866908 | 978554430 | 978323049 | 1018027765 | 484408 | GCaMP6f | Slc17a7-IRES2-Cre/wt;Camk2a-tTA/wt;Ai93(TITL-G... | [Slc17a7-IRES2-Cre, Camk2a-tTA] | Slc17a7-IRES2-Cre | Ai93(TITL-GCaMP6f) | M | ... | False | Novel >1 | 0 | 2 | 6 | 2019-11-08 08:52:05.141000+00:00 | MESO.1 | 2021-08-12 | 942701302 | 332 |
5 rows × 30 columns
Thats 7 different recording sessions for this single imaging plane. Remember that one ophys_container_id is linked to one imaging plane, recorded in multiple sessions
What are the session types for this container?¶
container_experiments.session_type.unique()
array(['OPHYS_1_images_G', 'OPHYS_2_images_G_passive', 'OPHYS_3_images_G',
'OPHYS_4_images_H', 'OPHYS_6_images_H'], dtype=object)
Reminder about structure & terminology of the dataset¶

Reminder about cre lines¶

Reminder about dataset variants aka project_codes¶

Reminder about session types¶


Identifying experiments and sessions of interest¶
Get all experiments for one container from an Sst-IRES-Cre mouse in the VisualBehaviorTask1B project code¶
# get all Sst experiments in the relevant project code
sst_experiments = ophys_experiments[(ophys_experiments.cre_line=='Sst-IRES-Cre')&
(ophys_experiments.project_code=='VisualBehaviorTask1B')]
# pick some container from this set
ophys_container_id = sst_experiments.ophys_container_id.unique()[1]
print(ophys_container_id)
941373529
# what experiments are there for this container?
sst_container_experiments = sst_experiments[sst_experiments.ophys_container_id==ophys_container_id]
sst_container_experiments
| behavior_session_id | ophys_session_id | ophys_container_id | mouse_id | indicator | full_genotype | driver_line | cre_line | reporter_line | sex | ... | passive | experience_level | prior_exposures_to_session_type | prior_exposures_to_image_set | prior_exposures_to_omissions | date_of_acquisition | equipment_name | published_at | isi_experiment_id | file_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ophys_experiment_id | |||||||||||||||||||||
| 955276580 | 955110047 | 954981981 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 20 | 0 | 2019-09-25 13:31:46+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 577 |
| 956903375 | 956213722 | 955991376 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | True | Familiar | 0 | 21 | 1 | 2019-09-26 11:13:06.028000+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 578 |
| 957652800 | 957331258 | 957189583 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Familiar | 0 | 22 | 2 | 2019-09-27 11:57:27.017000+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 579 |
| 959337347 | 958994511 | 958931715 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Novel >1 | 2 | 2 | 5 | 2019-10-01 11:20:24.445000+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 580 |
| 960351917 | 959893902 | 959751299 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | True | Novel >1 | 0 | 3 | 6 | 2019-10-02 11:46:04.405000+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 581 |
| 960960480 | 960697332 | 960593969 | 941373529 | 470784 | GCaMP6f | Sst-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt | [Sst-IRES-Cre] | Sst-IRES-Cre | Ai148(TIT2L-GC6f-ICL-tTA2) | F | ... | False | Novel >1 | 0 | 4 | 7 | 2019-10-03 11:25:31.172000+00:00 | CAM2P.5 | 2021-03-25 | 899181668 | 582 |
6 rows × 30 columns
Load the BehaviorOphysExperiment dataset for each ophys_experiment_id in the container and plot the max intensity projection - are they well aligned across sessions? can you identify the same neurons?¶
import matplotlib.pyplot as plt
# ophys_experiment_ids are the index of the ophys_experiment_table
ophys_experiment_ids = sst_container_experiments.index.values
# create figure axis
fig, ax = plt.subplots(1, len(ophys_experiment_ids), figsize=(20,5))
# enumerate over experiments in this container
for i, ophys_experiment_id in enumerate(ophys_experiment_ids):
# get the dataset object
dataset = cache.get_behavior_ophys_experiment(ophys_experiment_id=ophys_experiment_id)
# get the max intensity projection and plot on the appropriate axis
ax[i].imshow(dataset.max_projection.data, cmap='gray')
ax[i].set_title(ophys_experiment_id)
behavior_ophys_experiment_955276580.nwb: 100%|██████████| 313M/313M [00:13<00:00, 22.8MMB/s] behavior_ophys_experiment_956903375.nwb: 100%|██████████| 322M/322M [00:13<00:00, 23.4MMB/s] behavior_ophys_experiment_957652800.nwb: 100%|██████████| 318M/318M [00:13<00:00, 22.8MMB/s] behavior_ophys_experiment_959337347.nwb: 100%|██████████| 342M/342M [00:14<00:00, 22.8MMB/s] behavior_ophys_experiment_960351917.nwb: 100%|██████████| 327M/327M [00:14<00:00, 22.6MMB/s] behavior_ophys_experiment_960960480.nwb: 100%|██████████| 332M/332M [00:18<00:00, 18.0MMB/s]
Get all imaging planes recorded during one session with novel images in a Vip mouse imaged on Multiscope¶
# get all Vip sessions in the Multiscope project code
vip_sessions = ophys_sessions[(ophys_sessions.cre_line=='Vip-IRES-Cre')&
(ophys_sessions.project_code=='VisualBehaviorMultiscope')&
(ophys_sessions.prior_exposures_to_image_set==0)]
# ophys_session_id is the index of the ophys_session_table
ophys_session_id = vip_sessions.index.values[0]
# look at info for this ophys session
vip_sessions.loc[ophys_session_id]
behavior_session_id 1050620316 ophys_container_id [1046576772, 1046576775, 1046576784, 104657678... mouse_id 523922 indicator GCaMP6f full_genotype Vip-IRES-Cre/wt;Ai148(TIT2L-GC6f-ICL-tTA2)/wt driver_line [Vip-IRES-Cre] cre_line Vip-IRES-Cre reporter_line Ai148(TIT2L-GC6f-ICL-tTA2) sex M age_in_days 183 imaging_plane_group_count 4 project_code VisualBehaviorMultiscope session_type OPHYS_4_images_B session_number 4 image_set images_B behavior_type active_behavior experience_level Novel 1 prior_exposures_to_session_type 0 prior_exposures_to_image_set 0 prior_exposures_to_omissions 7 date_of_acquisition 2020-09-15 09:31:58.082000+00:00 equipment_name MESO.1 num_depths_per_area 4 ophys_experiment_id [1050762966, 1050762969, 1050762972, 105076297... num_targeted_structures 2 Name: 1050597678, dtype: object
Plot the average dF/F trace for each of the experiments in this session for a 5 minute time period¶
# get all the ophys_experiment_ids (corresponding to imaging planes) for this session
ophys_experiment_ids = vip_sessions.loc[ophys_session_id].ophys_experiment_id
print(ophys_experiment_ids)
[1050762966, 1050762969, 1050762972, 1050762974, 1050762975, 1050762977]
# create figure axis
fig, ax = plt.subplots(1,1, figsize=(15,4))
# enumerate over experiments in this session
for i, ophys_experiment_id in enumerate(ophys_experiment_ids):
# get the dataset object
dataset = cache.get_behavior_ophys_experiment(ophys_experiment_id=ophys_experiment_id)
# get ophys timestamps
ophys_timestamps = dataset.ophys_timestamps
# get the population average dF/F trace
dff_traces = dataset.dff_traces
# dff_traces is a dataframe with a column 'dff'
# get the values of this column and turn into a matrix of n_cells x timepoints
dff_traces = np.vstack(dff_traces.dff.values)
# take the mean over the cell axis
average_dFF = np.mean(dff_traces, axis=0)
# get the imaging_depth and targeted_structure for this experiment
imaging_depth = dataset.metadata['imaging_depth']
targeted_structure = dataset.metadata['targeted_structure']
# plot it, including the imaging_depth and targeted_structure in the legend label
ax.plot(ophys_timestamps, average_dFF, label=targeted_structure+'_'+str(imaging_depth))
ax.set_title(dataset.metadata['cre_line']+', ophys_session_id: '+str(ophys_session_id))
ax.set_ylabel('dF/F')
ax.set_xlabel('time (seconds)')
ax.set_xlim(5*60, 10*60)
ax.legend()
behavior_ophys_experiment_1050762966.nwb: 100%|██████████| 248M/248M [00:10<00:00, 22.6MMB/s] behavior_ophys_experiment_1050762969.nwb: 100%|██████████| 258M/258M [00:11<00:00, 23.3MMB/s] behavior_ophys_experiment_1050762972.nwb: 100%|██████████| 251M/251M [00:10<00:00, 25.1MMB/s] behavior_ophys_experiment_1050762974.nwb: 100%|██████████| 251M/251M [00:10<00:00, 23.6MMB/s] behavior_ophys_experiment_1050762975.nwb: 100%|██████████| 259M/259M [00:10<00:00, 25.0MMB/s] behavior_ophys_experiment_1050762977.nwb: 100%|██████████| 250M/250M [00:10<00:00, 23.9MMB/s]
<matplotlib.legend.Legend at 0x7faebb67d3d0>
Ophys Cells Table¶
cells_table = cache.get_ophys_cells_table()
cells_table.head()
| ophys_experiment_id | cell_specimen_id | |
|---|---|---|
| cell_roi_id | ||
| 1080884343 | 775614751 | 1086496928 |
| 1080884173 | 775614751 | 1086496914 |
| 1080883843 | 775614751 | 1086496838 |
| 1080886674 | 775614751 | 1086491756 |
| 1080885658 | 775614751 | 1086491699 |
How many cells per experiment?¶
cell_per_exp = cells_table.groupby('ophys_experiment_id').size()
fig = plt.hist(cell_per_exp, bins=50)
plt.xlabel('Cell count')
plt.ylabel('Number of experiments')
plt.show()
cell_per_exp.describe()
count 1936.000000 mean 68.732438 std 95.376045 min 1.000000 25% 11.000000 50% 22.000000 75% 99.000000 max 666.000000 dtype: float64
Merge the cell counts into the ophys experiments table
ophys_experiments['n_cells'] = ophys_experiments.index.map(cell_per_exp)
Now we can look at the cell count by depth, for example
fig, ax = plt.subplots(figsize=(30, 10))
ax.scatter(ophys_experiments['imaging_depth'], ophys_experiments['n_cells'], alpha=.3)
ax.set_xlabel('Imaging depth (microns)')
ax.set_ylabel('Cell count')
plt.show()
Or by cre-line
import seaborn as sns
sns.boxplot(data=ophys_experiments, x='n_cells', y='cre_line')
<Axes: xlabel='n_cells', ylabel='cre_line'>